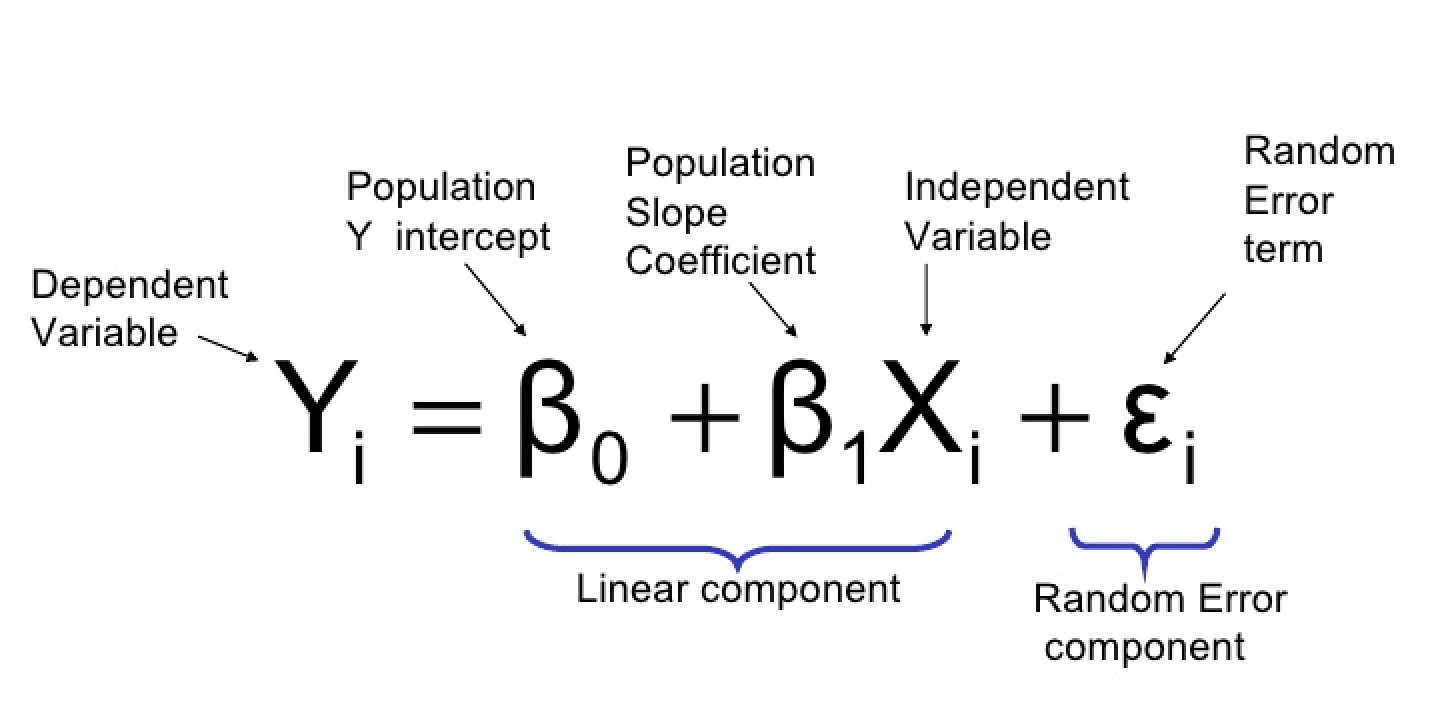
When we have a simple linear regression with two levels (a t-test) the predicted value is simply the overall mean value for that group. 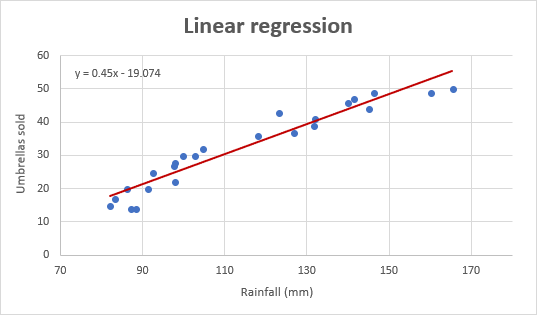
Recall, the residuals are the difference between actual/observed value and the predicted value. The question original question was about residuals from the t-test. The intercept coefficient is 5.006, which means that when versicolor is set to “0” in the model (0 * 0.93 = 0) all we are left with is the intercept, which is the mean value for setosa’s sepal length, the same as we saw in our t-test. This is the same result we obtained with our t-test above. Notice that the slope coefficient for versicolor is 0.93, indicating it’s sepal length is, on average, 0.93 greater than setosa’s sepal length. # add an intercept constant, since it isn't done automatically # Linear model to compare results with t-test (convert the species types of dummy variables) We fit our model using the statsmodels library. Now that we see what the output looks like, let’s confirm that this is indeed just linear regression! We also get the degrees of freedom, t-statistic, and p-value, along with several measures of effect size. We see the observed difference, versicolor has a sepal length 0.93 (5.006 – 5.936) longer that setosa, on average. We can see the summary stats for both groups at the top. To get a better look at the underlying comparison, I’ll instead fit the t-test using the researchpy library. They simply return the t-statistics, p-value, and degrees of freedom. Unfortunately, the output of both of these approaches leaves a lot to be desired. Stats.ttest_ind(a = df2 = 'setosa'],ī = df2 = 'versicolor'], Since a t-test is a comparison of means between two groups, I’ll create a data set with only the setosa and versicolor species.įirst I build the t-test in two common stats libraries in python, statsmodels and scipy. For this tutorial the variable we will look at is Sepal Length, which appears to different between Species. The Jupyter Notebook I’ve made available on GITHUB has a number of EDA steps. The data we will use is the iris data set, available in the numpy library. In this sense, a t-test is just a simple linear regression with a single categorical predictor (independent) variable that has two levels (e.g., Male & Female) while ANOVA is a simple linear regression with a single predictor variable that has more than two levels (e.g., Cat, Dog, Fish).Ĭomplete code is available on my GITHUB page. As such, an easier way of thinking about them is that they are a different way of looking at a regression output. The thing we need to remember about t-tests and ANOVA is that they are general linear models. The question was regarding how to get the residuals from a t-test. I had someone ask me a question the other day about t-tests.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
